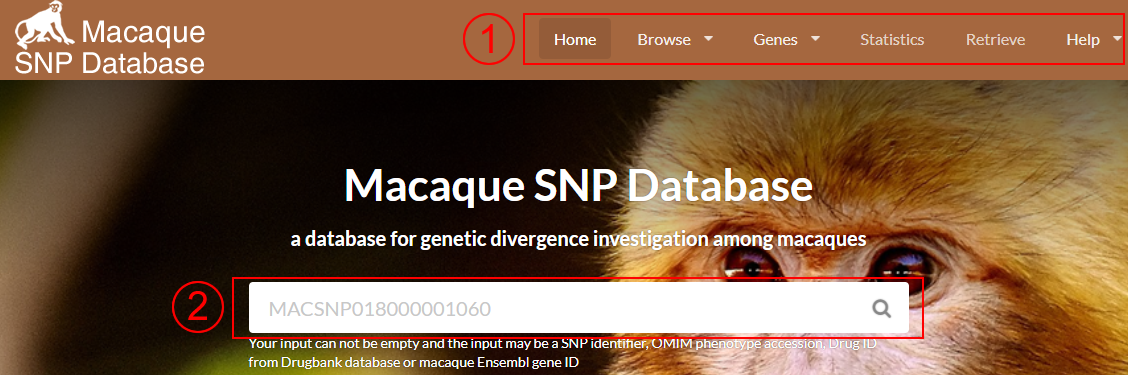
Quick search function allows you to provide a SNV identifier (individual, group-specific, species-specific or non-redundant SNV), macaque Ensembl gene ID, OMIM phenotype accession number or drug ID from Drugbank database to query item.
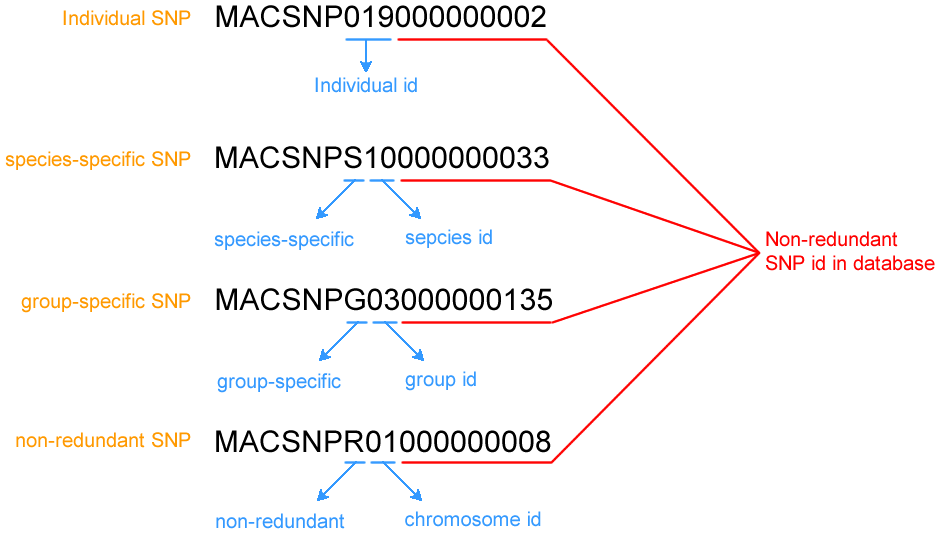
The database will automatically generate a unique identifier for individual, specific or non-redundant SNV. All the SNV identifier start with "MACSNV". The generation rule of identifers are shown in the following Figure.
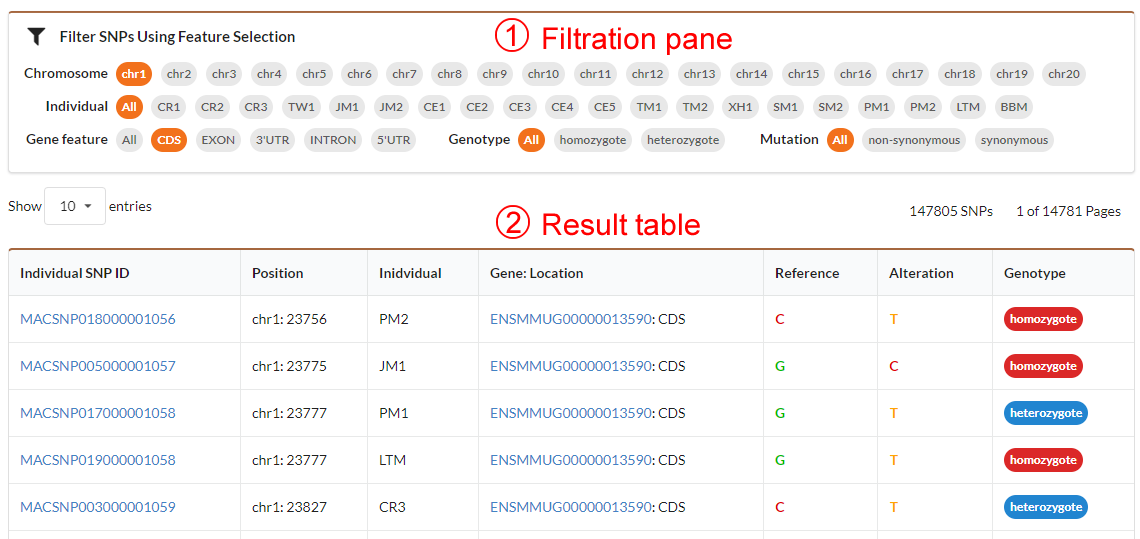
The individual SNVs browse page includes the filtration pane (1) with the SNV result table (2) located under it. Through filtration pane, you may filter SNVs in table by specifying chrosome number, individual, gene feature (CDS, UTR or Intron etc.), genotype (homozygote or heterozygote) and mutation type (non-synonymous or synonymous). The result table listed SNV information including ID, position in chromosome, reference, alteration, individual, gene location and genotype. By clicking the SNV ID hyperlink, you can obtain detailed information for SNV.
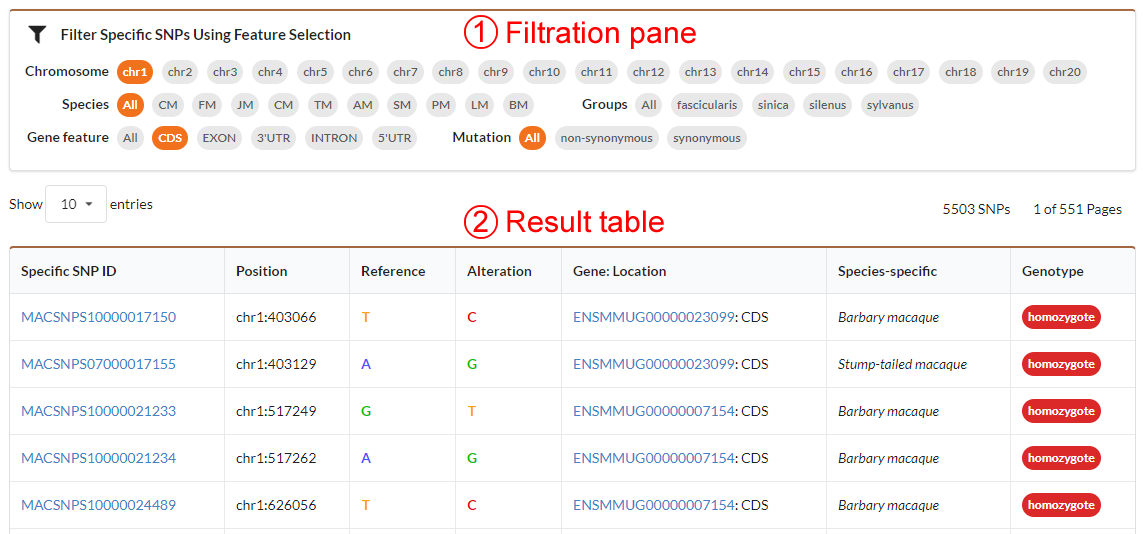
The specific SNVs browse page has the same interface with individual SNVs browe page. The filtration pane allows you to select a species or group to get a list of corresponding specific SNVs. The detailed information for specific SNV can be obtained by clicking the specific SNV ID hyperlink.
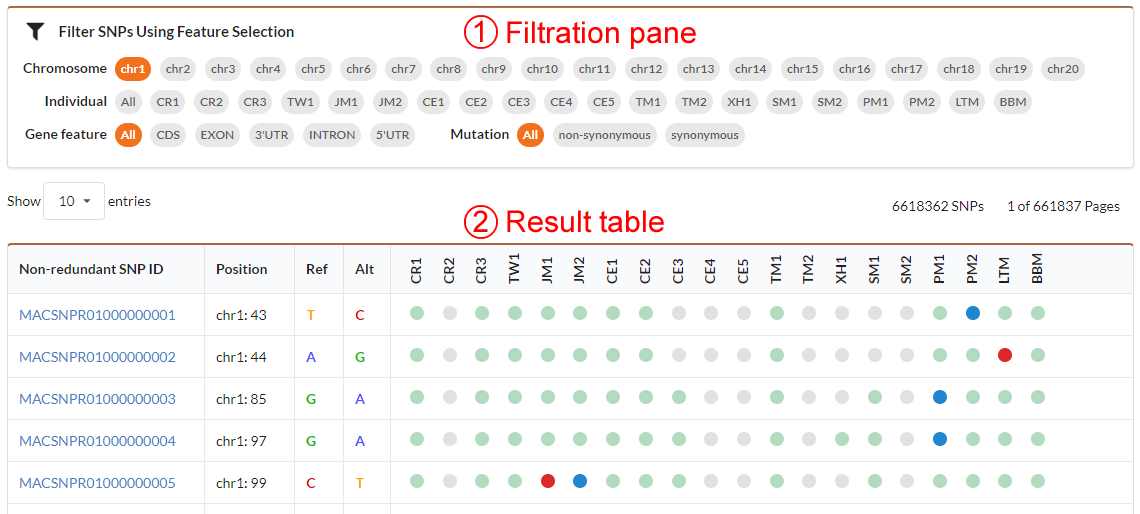
In the non-redundant SNVs browse page, in addition to non-redundant SNV information, you also get a genotyping comparative map of individuals. The genotyes are indicated by colored circle dots (red: homozygote, blue: heterozygote, green: not variant for that individual, gray: no reads mapped to this position for that individual). Users can select several individuals in filtration pane to generate a genotyping comparative map. Users may click the non-redundant SNV ID hyperlink to obtain detail information or click the red or blue dot to view individual SNV.
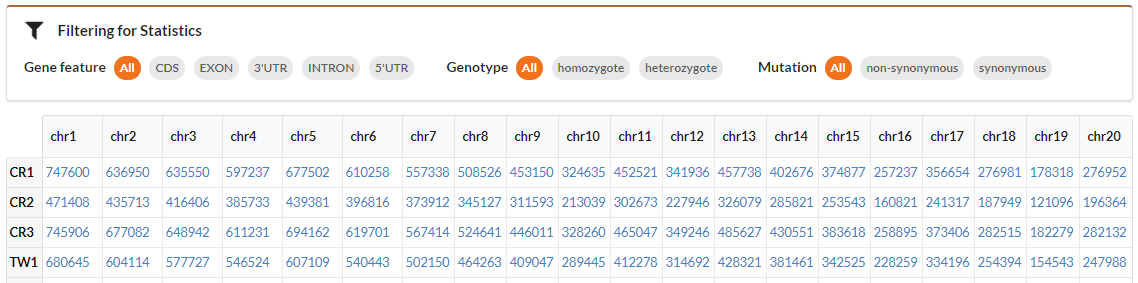
MACSNVdb summarizes SNV number by chromosome for each individuals and generates a statistical table shown in above figure. Each column indicates the SNV number in a chromosome of each individuals and Each row indicates the SNV number of a individual in each chromosome. Click the number, you can view the SNVs in a given chromosome and individual. You can use filters in filtration pane to get SNV summary table in gene region. You can also restrict the genotype and mutation type.
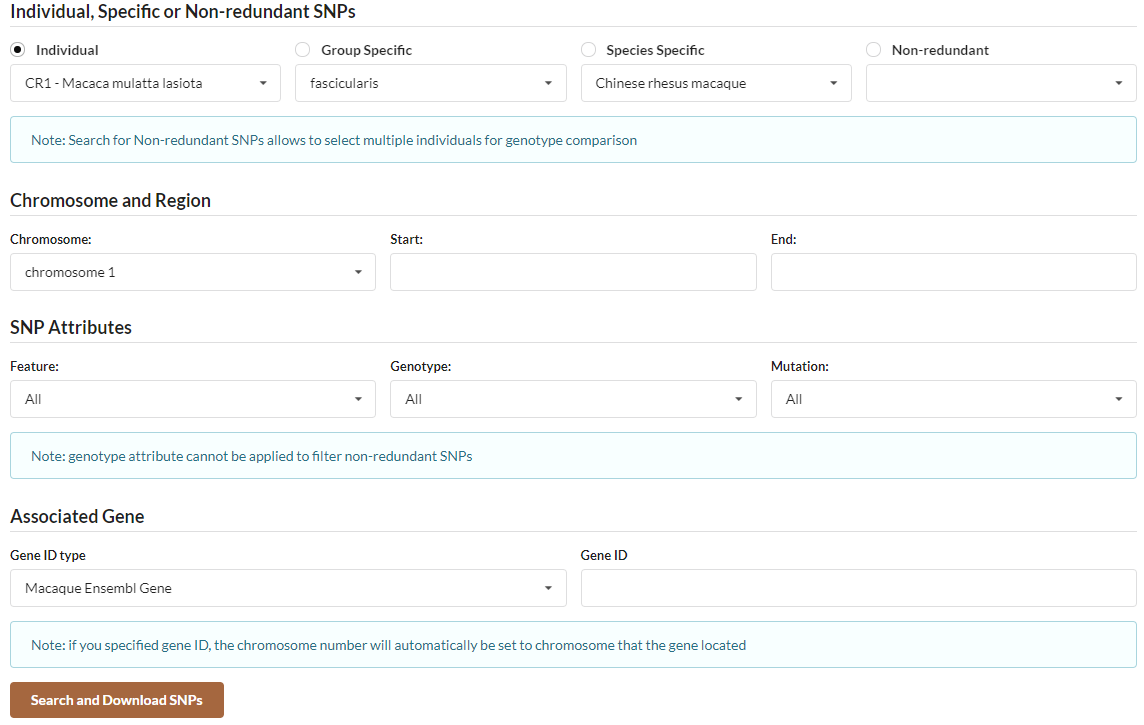
MACSNVdb provide a flexible search function for you to retrieve desired SNVs. The above figure shows the interface of search page. You can select to search individual, group-specific, species-specific or non-redundant SNVs. You can set chromosome number, start position and end position. You can also set location of SNVs in gene, genotype and mutation type to restrict search result. Centainly, you can specify gene id to get gene-associated SNVs. When you search non-redundant SNVs, you can select multiple individuals to genotype comparison among individuals.
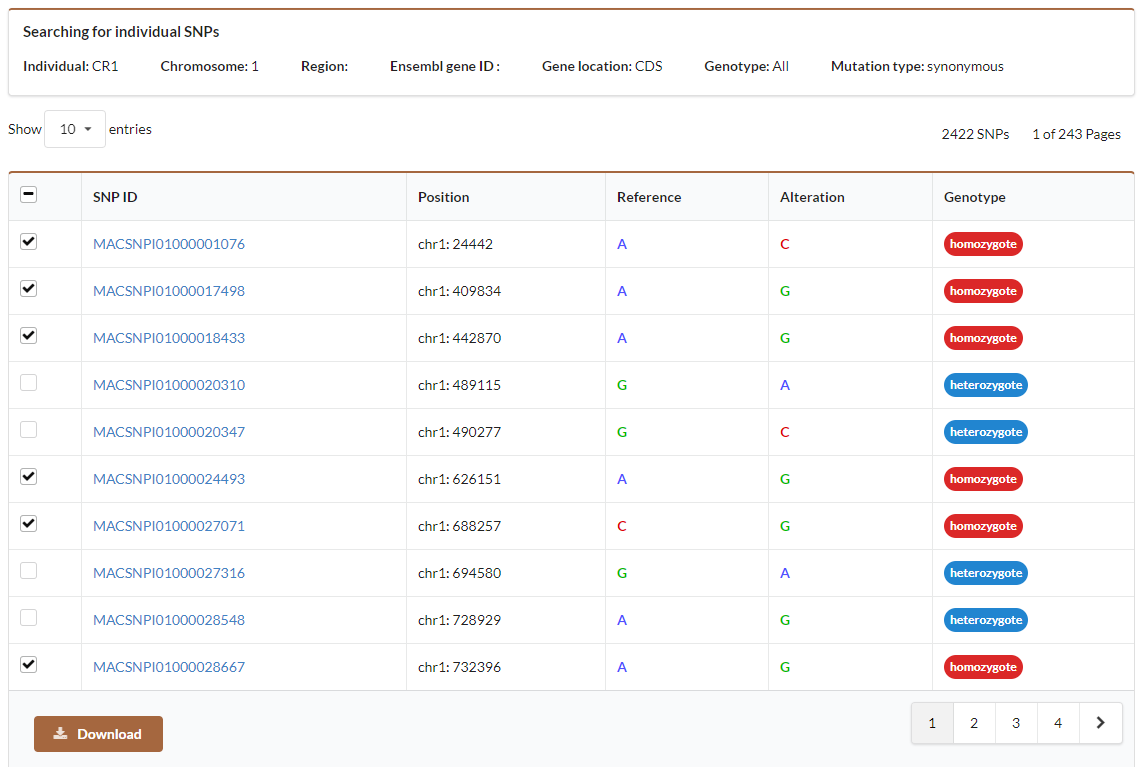
The above figure shows the search result of individual SNVs. You can select SNVs that you are interested and then click download button to obtain detailed information of selected SNVs.